RS/tRNA Foundational Publication Support
Amiram, Miriam, Adrian D. Haimovich, Chenguang Fan, Yane-Shih Wang, Hans-Rudolf Aerni, Ioanna Ntai, Daniel W. Moonan, et al. (dec) 2015. “Evolution Of Translation Machinery In Recoded Bacteria Enables Multi-Site Incorporation Of Nonstandard Amino Acids”. Nature Biotechnology 33: 1272-1279. doi:10.1038/nbt.3372.
Wang, Lei, Zhiwen Zhang, Ansgar Brock, and Peter G Schultz. (2003) 2003. “Addition Of The Keto Functional Group To The Genetic Code Of Escherichia Coli.”. Proceedings Of The National Academy Of Sciences Of The United States Of America 100 (1): 56-61.
Young, Travis S, Insha Ahmad, Jun A Yin, and Peter G Schultz. (2010) 2010. “An Enhanced System For Unnatural Amino Acid Mutagenesis In E. Coli.”. Journal Of Molecular Biology 395 (2): 361-74. doi:10.1016/j.jmb.2009.10.030.
Young, Douglas D, Travis S Young, Michael Jahnz, Insha Ahmad, Glen Spraggon, and Peter G Schultz. (2011) 2011. “An Evolved Aminoacyl-Trna Synthetase With Atypical Polysubstrate Specificity.”. Biochemistry 50 (11): 1894-900. doi:10.1021/bi101929e.
RS/tRNA Usage Publications
Arsiwala, Ammar, Chad Varner, Jessica N McCaffery, Andrew Kell, Geetanjali Pendyala, Ana Castro, Vivek Hariharan, Alberto Moreno, and Ravi S Kane. (2019) 2019. “Nanopatterning Protein Antigens To Refocus The Immune Response.”. Nanoscale 11 (32): 15307-15311. doi:10.1039/c9nr05145g.
Castro, Ana, Juan Manuel Carreño, James Duehr, Florian Krammer, and Ravi S. Kane. (aug) 2021. “Refocusing The Immune Response To Selected Epitopes On A Zika Virus Protein Antigen By Nanopatterning”. Advanced Healthcare Materials 10: e2002140. doi:10.1002/adhm.202002140.
RS/tRNA Pair Development Year
2015
ncAA(s) Incorporated
p-acetyl-L-phenylalanine
ncAA Structure (png, jpg, jpeg)
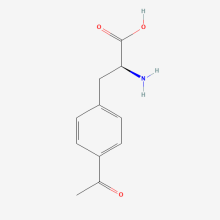
ncAA Utility
Site-directed spin labeling, electron paramagnetic resonance
O-tert-butyl-L-tyrosine (Tby)
ncAA Structure (png, jpg, jpeg)

ncAA Utility
NMR tag for proteins
p-iodo-L-phenylalanine
ncAA Structure (png, jpg, jpeg)

ncAA Utility
Can be used for phasing in protein crystallography.
Trifluoromethyl-L phenylalanine
ncAA Structure (png, jpg, jpeg)

ncAA Utility
19F Probe for protein
NMR
NMR
O-methyl-L-tyrosine
ncAA Structure (png, jpg, jpeg)

ncAA Utility
Labelling of muscles
(global)
(global)
RS Organism of Origin
Parent RS
RS Mutations
Y32L
L65V
D158G
I159C
L162R
A167D
R257G
L65V
D158G
I159C
L162R
A167D
R257G
tRNA Organism of Origin
Parent tRNA
tRNA Anticodon
CUA
Other tRNA Mutations
C17A
U17aG
U20C
G37A
U47G
U17aG
U20C
G37A
U47G
RS/tRNA Availability
Addgene Plasmid #73544
RS/tRNA Additional Notes
This RS/tRNA pair was evolved in the C321.A E. coli strain from the AcetylPhe RS (pAcF-RS) originally reported in in 2002 foundational paper, used as a model RS in the 2010 foundational paper and characterized for permissivity in the 2011 foundational paper. In 2015 further residues were allowed to evolve to optimize p-acetyl-Phe and tRNA interactions and improve expression of multiple ncAA containing proteins using low RS levels. This RS showed a 15-fold increased expression of GFP(3UAG) compared to pAcFRS using a chromosome incorporated RS. Figure 4a of foundational paper shows that when included as multicopy plasmid it still highly outperforms the starting RS for incorporating 30 ncAAs into a single protein construct, but does not outperform the original unevolved RS when expressing GFP with 3 ncAAs, and it also has a lower fidelity (higher protein production in the absence of ncAA). This RS was also shown to be highly promiscuous, incorporating Phe variants pAcetyl, pIodo, pBromo, pChloro, pMethyl, ptrifluoromethyl along with methylTyr, 2-napthylAla and styrylAla. Was selected to discrimate against pAzf and did when chromosome incorporated, but not when provided as a multicopy plasmid. Also especially effective at incorporating O-methyl-Tyr, p-IodoPhe, p-trifluormethylPhe, p-tert-butyl-Phe, O-tert-Butyl and p-phenylPhe.