RS/tRNA Foundational Publication Support
Truong, Frank, Tae Hyeon Yoo, Thomas Lampo, and David A Tirrell. (2012) 2012. “Two-Strain, Cell-Selective Protein Labeling In Mixed Bacterial Cultures.”. Journal Of The American Chemical Society 134 (20): 8551-6. doi:10.1021/ja3004667.
Marchand, J, M Neugebauer, M Ing, C-I Lin, J Pelton, and M C Y Chang. (2019) 2019. “Discovery Of A Pathway For Terminal-Alkyne Amino Acid Biosynthesis.”. Nature 567 (7748): 420-424. doi:10.1038/s41586-019-1020-y.
RS/tRNA Pair Development Year
2012
ncAA(s) Incorporated
propargylglycine
ncAA Structure (png, jpg, jpeg)
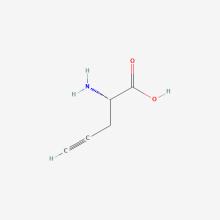
ncAA Utility
copper-catalyzed alkyne-azide click chemistry (CuACC) can be used for time-resolved, cell-selective proteomic analyses
RS Organism of Origin
Parent RS
RS Mutations
L13P
A256G
P257T
Y260Q
H301F
A331V
A256G
P257T
Y260Q
H301F
A331V
tRNA Organism of Origin
Parent tRNA
tRNA Anticodon
CAU
Other tRNA Mutations
none
RS/tRNA Availability
n/a
RS/tRNA Additional Notes
The RS/tRNA pair incorporates propargylglycine (Pra) at all Met positions with near 100% replacement of Met in a Met auxotroph (E coli strain TYJV2) with Met-depleted M9 minimal medium supplemented with 4 mM Pra. Incorporation was at ~10% level in M9 medium supplemented with 269 µM Met and 4 mM Pra. This RS does not incorporate azidonorleucine (AnL), so when it is used in one cell type in a mixed culture and the NLL-MetRS in another, it allows mixed cell types to be selectively labeled using Pra and AnL, which can then via click chemistry (first AnL and then Pra) be independently be ligated for analyses.