RS/tRNA Foundational Publication Support
Rogerson, D. T., A. Sachdeva, K. Wang, T. Haq, A. Kazlauskaite, S. M. Hancock, N. Huguenin-Dezot, M. M. K. Muqit, A. M. Fry, and R. Bayliss. n.d. “Efficient Genetic Encoding Of Phosphoserine And Its Nonhydrolyzable Analog” 11: 496+.
Beránek, V., C. D. Reinkemeier, M. S. Zhang, A. D. Liang, G. Kym, and J. W. Chin. n.d. “Genetically Encoded Protein Phosphorylation In Mammalian Cells” 25: 1067+.
Zhu, Phillip, Philip R Gafken, Ryan A Mehl, and Richard B Cooley. (2019) 2019. “A Highly Versatile Expression System For The Production Of Multiply Phosphorylated Proteins.”. Acs Chemical Biology 14 (7): 1564-1572. doi:10.1021/acschembio.9b00307.
Vesely, Cat Hoang, Patrick N Reardon, Zhen Yu, Elisar Barbar, Ryan A Mehl, and Richard B Cooley. (2022) 2022. “Accessing Isotopically Labeled Proteins Containing Genetically Encoded Phosphoserine For Nmr With Optimized Expression Conditions.”. The Journal Of Biological Chemistry 298 (12): 102613. doi:10.1016/j.jbc.2022.102613.
Zhu, Phillip, Stanislau Stanisheuski, Rachel Franklin, Amber Vogel, Cat Hoang Vesely, Patrick Reardon, Nikolai N Sluchanko, et al. (2023) 2023. “Autonomous Synthesis Of Functional, Permanently Phosphorylated Proteins For Defining The Interactome Of Monomeric 14-3-3Ζ.”. Acs Central Science 9 (4): 816-835. doi:10.1021/acscentsci.3c00191.
RS/tRNA Protocols and Structural Information
Zhu, Phillip, Ryan A Mehl, and Richard B Cooley. (2022) 2022. “Site-Specific Incorporation Of Phosphoserine Into Recombinant Proteins In .”. Bio-Protocol 12 (21). doi:10.21769/BioProtoc.4541.
Zhu, Phillip, Rachel Franklin, Amber Vogel, Stanislau Stanisheuski, Patrick Reardon, Nikolai N Sluchanko, Joseph S Beckman, Andrew Karplus, Ryan A Mehl, and Richard B Cooley. (2021) 2021. “Permaphos : Autonomous Synthesis Of Functional, Permanently Phosphorylated Proteins.”. Biorxiv : The Preprint Server For Biology. doi:10.1101/2021.10.22.465468.
RS/tRNA Usage Publications
Stuber, Katrin, Tobias Schneider, Jill Werner, Michael Kovermann, Andreas Marx, and Martin Scheffner. (2021) 2021. “Structural And Functional Consequences Of Nedd8 Phosphorylation.”. Nature Communications 12 (1): 5939. doi:10.1038/s41467-021-26189-9.
Huguenin-Dezot, Nicolas, Virginia De Cesare, Julien Peltier, Axel Knebel, Yosua Adi Kristaryianto, Daniel T Rogerson, Yogesh Kulathu, Matthias Trost, and Jason W Chin. (2016) 2016. “Synthesis Of Isomeric Phosphoubiquitin Chains Reveals That Phosphorylation Controls Deubiquitinase Activity And Specificity.”. Cell Reports 16 (4): 1180-1193. doi:10.1016/j.celrep.2016.06.064.
Chadda, Rahul, Vikas Kaushik, Iram Ahmad, Jaigeeth Deveryshetty, Alex Holehouse, Snorri Sigurdsson, Gargi Biswas, et al. (2024) 2024. “Partial Wrapping Of Single-Stranded Dna By Replication Protein A And Modulation Through Phosphorylation.”. Nucleic Acids Research 52 (19): 11626-11640. doi:10.1093/nar/gkae584.
RS/tRNA Pair Development Year
2015
ncAA(s) Incorporated
phosphoserine
ncAA Structure (png, jpg, jpeg)
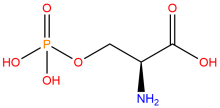
ncAA Utility
Used to produce proteins site-specifically phosphorylated at serine residues, typically at positions where phosphoserine naturally occurs as a post-translational modification
2-amino-4-phosphonobutanoic acid (nhpSer)
ncAA Structure (png, jpg, jpeg)
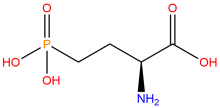
ncAA Utility
Used to produce proteins that mimic site-specific phosphorylation at a serine residue - where the phosphate group cannot by hydrolyzed - via genetic code expansion versus other protein phosphorylation strategies.
RS Organism of Origin
Parent RS
RS Mutations
E412P
E414F
P495M
I496W
F529S
E414F
P495M
I496W
F529S
tRNA Organism of Origin
Parent tRNA
tRNA Anticodon
CUA
Other tRNA Mutations
G29A
A31U
G34C
C35
G37A
U39A
C40U
C41U
A31U
G34C
C35
G37A
U39A
C40U
C41U
Multiple tRNAs?
Can also work with Sep-tRNA-v2 (developed in 2019).
This tRNA was shown in 2022 foundational paper to be preferable for pSer expressions done in minimal media, for instance, for isotopic labeling for NMR studies.
It was also in 2023 chosen as the RNA for the Permaphos GCE system to install nhpSer (system described separately under SepRS2/Sep-tRNA-v2 pair)
This tRNA was shown in 2022 foundational paper to be preferable for pSer expressions done in minimal media, for instance, for isotopic labeling for NMR studies.
It was also in 2023 chosen as the RNA for the Permaphos GCE system to install nhpSer (system described separately under SepRS2/Sep-tRNA-v2 pair)
RS/tRNA Availability
Component Addgene IDs are #173897 for SepRS2/Sep-tRNA(B4) machinery, and #174075 and #174076 for sfGFP WT and 150TAG constructs, respectively.
E. coli strains:
BL21(DE3) ΔserB - Addgene #34929
B95(DE3) ΔA ΔfabR ΔserB - Addgene #197655
Keio ΔserC - Obtained from GE-Dharmacon
The sfGFP/POI plasmids are not the ones originally used with this system, but work well with it. You can replace the sfGFP constructs by cloning in another POI.
E. coli strains:
BL21(DE3) ΔserB - Addgene #34929
B95(DE3) ΔA ΔfabR ΔserB - Addgene #197655
Keio ΔserC - Obtained from GE-Dharmacon
The sfGFP/POI plasmids are not the ones originally used with this system, but work well with it. You can replace the sfGFP constructs by cloning in another POI.
Used in what cell line?
RS/tRNA Additional Notes
2015 paper reports improved RS/tRNA pair which when used with EF-Sep led to full-fidelity incorporation of pSer into GST-calmodulin with little truncation, and myoglobin, ubiquitin (3 mg/L) and Nek7 (2 mg/L). Also shows that in an E coli (ΔserC + overexpressed SerB) strain the system gives high-fidelity of nhpSer in a myoglobin test protein.
2019 paper improves performance by putting the tools into the healthy, RF1-free B95(DE3) ΔA ΔfabR ΔserB E coli strain to create the "pSer-3.1G" system that can give truncation-free, full-fidelity incorporation at multiple sites. Produced 400 and 200 mg/L singly and double phosphorylated GFP, a quintuply phosphorylated GFP at 40 mg/L, and also efficiently produced doubly phosphorylated STING and triply phoshorylated BAD at over 5 mg/L.
2018 paper reports this RS/tRNA pair can lead to pSer incorporation in HEK293 cells with a knocked out Ser-phosphatase (PSPH), a mutated EF-1alpha (EF-1alpha-Sep),and an eRF1 E55D mutant. However incorporated pSer is readily hydrolyzed. Also in cells with overexpressed PSPH, knocked out PSAT (pSer amino transferase), EF-1alpha-Sep,and eRF1 E55D, pair could incorporate nhpSer at ~50% fidelity when cells were supplemented with exogenous ~10 mM nhpSer.
2019 paper improves performance by putting the tools into the healthy, RF1-free B95(DE3) ΔA ΔfabR ΔserB E coli strain to create the "pSer-3.1G" system that can give truncation-free, full-fidelity incorporation at multiple sites. Produced 400 and 200 mg/L singly and double phosphorylated GFP, a quintuply phosphorylated GFP at 40 mg/L, and also efficiently produced doubly phosphorylated STING and triply phoshorylated BAD at over 5 mg/L.
2018 paper reports this RS/tRNA pair can lead to pSer incorporation in HEK293 cells with a knocked out Ser-phosphatase (PSPH), a mutated EF-1alpha (EF-1alpha-Sep),and an eRF1 E55D mutant. However incorporated pSer is readily hydrolyzed. Also in cells with overexpressed PSPH, knocked out PSAT (pSer amino transferase), EF-1alpha-Sep,and eRF1 E55D, pair could incorporate nhpSer at ~50% fidelity when cells were supplemented with exogenous ~10 mM nhpSer.