RS/tRNA Foundational Publication Support
Lee, Sangsik, Seunghee Oh, Aerin Yang, Jihyo Kim, Dieter Söll, Daeyoup Lee, and Hee-Sung Park. 2013. “A Facile Strategy For Selective Incorporation Of Phosphoserine Into Histones”. Angewandte Chemie International Edition 52: 5771-5775. doi:10.1002/anie.201300531.
Pirman, Natasha L., Karl W. Barber, Hans R. Aerni, Natalie J. Ma, Adrian D. Haimovich, Svetlana Rogulina, Farren J. Isaacs, and Jesse Rinehart. (sep) 2015. “A Flexible Codon In Genomically Recoded Escherichia Coli Permits Programmable Protein Phosphorylation”. Nature Communications 6: 8130. doi:10.1038/ncomms9130.
Mohler, Kyle, Jack M Moen, Svetlana Rogulina, and Jesse Rinehart. (2023) 2023. “System-Wide Optimization Of An Orthogonal Translation System With Enhanced Biological Tolerance.”. Molecular Systems Biology 19 (8): e10591. doi:10.15252/msb.202110591.
RS/tRNA Protocols and Structural Information
Mohler, Kyle, and Jesse Rinehart. 2019. “Expression Of Authentic Post-Translationally Modified Proteins In Organisms With Expanded Genetic Codes”. Methods In Enzymology 626: 539-559. doi:10.1016/bs.mie.2019.07.017.
Gassaway, Brandon M, Jiaming Li, Ramin Rad, Julian Mintseris, Kyle Mohler, Tyler Levy, Mike Aguiar, et al. (2022) 2022. “A Multi-Purpose, Regenerable, Proteome-Scale, Human Phosphoserine Resource For Phosphoproteomics.”. Nature Methods 19 (11): 1371-1375. doi:10.1038/s41592-022-01638-5.
RS/tRNA Usage Publications
Perez-Pepe, Marcelo, Anthony W. Desotell, Hengyi Li, Wenxue Li, Bing Han, Qishan Lin, Daryl E. Klein, Yansheng Liu, Hani Goodarzi, and Claudio R. Alarcón. (may) 2023. “7Sk Methylation By Mettl3 Promotes Transcriptional Activity”. Science Advances 9: eade7500. doi:10.1126/sciadv.ade7500.
Di Mattia, Thomas, Arthur Martinet, Souade Ikhlef, Alastair G. McEwen, Yves Nominé, Corinne Wendling, Pierre Poussin-Courmontagne, et al. (dec) 2020. “Ffat Motif Phosphorylation Controls Formation And Lipid Transfer Function Of Inter-Organelle Contacts”. The Embo Journal 39: e104369. doi:10.15252/embj.2019104369.
Palani, Saravanan, Darius Koester, and Mohan K. Balasubramanian. 2020. “Phosphoregulation Of Tropomyosin-Actin Interaction Revealed Using A Genetic Code Expansion Strategy”. Wellcome Open Research 5: 161. doi:10.12688/wellcomeopenres.16082.1.
Baliova, Martina, and Frantisek Jursky. (aug) 2020. “Phosphorylation Of Serine 157 Protects The Rat Glycine Transporter Glyt2 From Calpain Cleavage”. Journal Of Molecular Neuroscience: Mn 70: 1216-1224. doi:10.1007/s12031-020-01529-4.
Mohler, Kyle, and Jesse Rinehart. 2019. “Expression Of Authentic Post-Translationally Modified Proteins In Organisms With Expanded Genetic Codes”. Methods In Enzymology 626: 539-559. doi:10.1016/bs.mie.2019.07.017.
Zheng, Huayu, Jingxuan He, Jinghui Li, Jing Yang, Martin L. Kirk, Linda J. Roman, and Changjian Feng. (feb) 2019. “Generation And Characterization Of Functional Phosphoserine-Incorporated Neuronal Nitric Oxide Synthase Holoenzyme”. Journal Of Biological Inorganic Chemistry: Jbic: A Publication Of The Society Of Biological Inorganic Chemistry 24: 1-9. doi:10.1007/s00775-018-1621-1.
Niemi, Natalie M., Gary M. Wilson, Katherine A. Overmyer, F.-Nora Vögtle, Lisa Myketin, Danielle C. Lohman, Kathryn L. Schueler, et al. (jul) 2019. “Pptc7 Is An Essential Phosphatase For Promoting Mammalian Mitochondrial Metabolism And Biogenesis”. Nature Communications 10: 3197. doi:10.1038/s41467-019-11047-6.
RS/tRNA Pair Development Year
2015
ncAA(s) Incorporated
phosphoserine
ncAA Structure (png, jpg, jpeg)
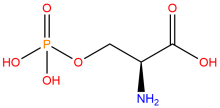
ncAA Utility
Used to produce proteins site-specifically phosphorylated at serine residues, typically at positions where phosphoserine naturally occurs as a post-translational modification
RS Organism of Origin
Parent RS
RS Mutations
E412S
E414I
P495R
I496R
K356E
N361D
L590I
E414I
P495R
I496R
K356E
N361D
L590I
tRNA Organism of Origin
Parent tRNA
tRNA Anticodon
CUA
Other tRNA Mutations
C20U
Multiple tRNAs?
In 2015, the G37A mutation was added to tRNA-Sep to make tRNA-SepA37 with improved UAG read-through. It more than doubled protein production.
In 2023, In addition to having A37, the C2:G71 base pair was changed to G2:C71 to decrease mischarging by GlyRS and ThrRS. This was called "tRNA-pSer-Opt"
In 2023, In addition to having A37, the C2:G71 base pair was changed to G2:C71 to decrease mischarging by GlyRS and ThrRS. This was called "tRNA-pSer-Opt"
RS/tRNA Availability
SepRS9, EF-Sep21 and tRNA-SepA37 are all on AddGene #68292. There are 4 copies of the tRNA on this plasmid, which may cause instability when propagating the plasmid.
The 2023 updated system used for studying peptides of the human pSer proteome expressed in E coli is Addgene #188537, and the rEcoli.XpS strain is Addgene #192872. This system expresses only 1 copy of the tRNA.
The 2023 updated system used for studying peptides of the human pSer proteome expressed in E coli is Addgene #188537, and the rEcoli.XpS strain is Addgene #192872. This system expresses only 1 copy of the tRNA.
Used in what cell line?
RS/tRNA Additional Notes
The EF-TU (EF-Sep21) has mutations H67R, E216V, D217G, F219Y, T229S, N274W to work with pSer. This lamba system yield 9-fold more GFP(site17) than the previous SepOTSα (SepRS, tRNA-Sep, EF-Sep) using C321.ΔA. Furthermore, the fidelity Sep was ~85% compared with ~50%. Also made MEK1 with pSer at sites 218 and 222 at 2 mg/L, but due to near-cognate suppression, the fidelity was only ~30% based on phostag gel analysis.