RS/tRNA Foundational Publication Support
Liu, Chang C, and Peter G Schultz. (2006) 2006. “Recombinant Expression Of Selectively Sulfated Proteins In Escherichia Coli.”. Nature Biotechnology 24 (11): 1436-40.
Schwessinger, Benjamin, Xiang Li, Thomas L Ellinghaus, Leanne Jade G Chan, Tong Wei, Anna Joe, Nicholas Thomas, et al. (2016) 2016. “A Second-Generation Expression System For Tyrosine-Sulfated Proteins And Its Application In Crop Protection.”. Integrative Biology : Quantitative Biosciences From Nano To Macro 8 (4): 542-5. doi:10.1039/c5ib00232j.
Chen, Yuda, Shikai Jin, Mengxi Zhang, Yu Hu, Kuan-Lin Wu, Anna Chung, Shichao Wang, et al. (2022) 2022. “Unleashing The Potential Of Noncanonical Amino Acid Biosynthesis To Create Cells With Precision Tyrosine Sulfation.”. Nature Communications 13 (1): 5434. doi:10.1038/s41467-022-33111-4.
RS/tRNA Protocols and Structural Information
Liu, Chang C, Susan E Cellitti, Bernhard H Geierstanger, and Peter G Schultz. (2009) 2009. “Efficient Expression Of Tyrosine-Sulfated Proteins In E. Coli Using An Expanded Genetic Code.”. Nature Protocols 4 (12): 1784-9. doi:10.1038/nprot.2009.188.
RS/tRNA Usage Publications
Liu, Chang C, Antha Mack V, Meng-Lin Tsao, Jeremy H Mills, Hyun Soo Lee, Hyeryun Choe, Michael Farzan, Peter G Schultz, and Vaughn Smider V. (2008) 2008. “Protein Evolution With An Expanded Genetic Code.”. Proceedings Of The National Academy Of Sciences Of The United States Of America 105 (46): 17688-93. doi:10.1073/pnas.0809543105.
Liu, Chang C, Hyeryun Choe, Michael Farzan, Vaughn Smider V, and Peter G Schultz. (2009) 2009. “Mutagenesis And Evolution Of Sulfated Antibodies Using An Expanded Genetic Code.”. Biochemistry 48 (37): 8891-8. doi:10.1021/bi9011429.
Schwessinger, Benjamin, Xiang Li, Thomas L Ellinghaus, Leanne Jade G Chan, Tong Wei, Anna Joe, Nicholas Thomas, et al. (2016) 2016. “A Second-Generation Expression System For Tyrosine-Sulfated Proteins And Its Application In Crop Protection.”. Integrative Biology : Quantitative Biosciences From Nano To Macro 8 (4): 542-5. doi:10.1039/c5ib00232j.
RS/tRNA Pair Development Year
2006
ncAA(s) Incorporated
sulfo-L-tyrosine
ncAA Structure (png, jpg, jpeg)
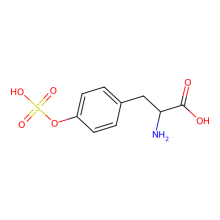
ncAA Utility
Labelling of muscles (global)
Dipeptide (DanAla-‐Ala) required for uptake and bioavailability
Dipeptide (DanAla-‐Ala) required for uptake and bioavailability
RS Organism of Origin
Parent RS
RS Mutations
Y32L
L65P
D158G
I159C
L162K
L65P
D158G
I159C
L162K
tRNA Organism of Origin
Parent tRNA
tRNA Anticodon
CUA
Other tRNA Mutations
C17A
U17aG
U20C
G37A
U47G
U17aG
U20C
G37A
U47G
RS/tRNA Availability
The RS is available in AddGene Plasmid #82417 (in the pULTRA plasmid system as shown in the second foundational publication)
The enzymes needed to make sulfoTyr endogenously in E. coli are encoded in Addgene #188983 (L-arabinose inducable)
The enzymes needed to make sulfoTyr endogenously in E. coli are encoded in Addgene #188983 (L-arabinose inducable)
Used in what cell line?
RS/tRNA Additional Notes
In DH10B cells, with 2 mM sulfoTyr, gave high fidelity incorporation into Z-domain(7), noting that some (<10%) of the sulfo-group was lost during MS analysis. And with 10 mM sulfoTyr, yielded hirudin(63) at 5mg/L compared to WT yield of 12 mg/L. In the pULTRA plasmid yielded ~75% efficient expression of GFP with one TAG site and ~25% efficient expression of GFP with three TAG sites, and high fidelity using the RF1- C321.ΔA cell line.
The Chen et al. 2022 foundational paper reports how sulfo-Tyr can also be biosynthesized using the plasmid pEvol-NnSULT1C1-cysDNCQ (Addgene #188983) in the BW25113 ΔcysH E. coli strain, providing ~10-fold improvement in sfGFP-TAG expression compared to supplementing media with 1 mM sTyr.
The Chen et al. 2022 foundational paper reports how sulfo-Tyr can also be biosynthesized using the plasmid pEvol-NnSULT1C1-cysDNCQ (Addgene #188983) in the BW25113 ΔcysH E. coli strain, providing ~10-fold improvement in sfGFP-TAG expression compared to supplementing media with 1 mM sTyr.