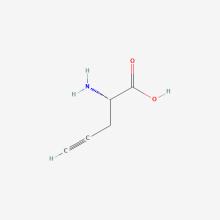
ncAA Structure (png, jpg, jpeg)
ncAA Foundational Publication Support
Truong, Frank, Tae Hyeon Yoo, Thomas Lampo, and David A Tirrell. (2012) 2012. “Two-Strain, Cell-Selective Protein Labeling In Mixed Bacterial Cultures.”. Journal Of The American Chemical Society 134 (20): 8551-6. doi:10.1021/ja3004667.
Marchand, J, M Neugebauer, M Ing, C-I Lin, J Pelton, and M C Y Chang. (2019) 2019. “Discovery Of A Pathway For Terminal-Alkyne Amino Acid Biosynthesis.”. Nature 567 (7748): 420-424. doi:10.1038/s41586-019-1020-y.
ncAA Usage Publications
Lang, Kathrin. (2019) 2019. “Installing Terminal-Alkyne Reactivity Into Proteins In Engineered Bacteria.”. Biochemistry 58 (24): 2703-2705. doi:10.1021/acs.biochem.9b00392.
ncAA Utility
copper-catalyzed alkyne-azide click chemistry (CuACC) can be used for time-resolved, cell-selective proteomic analyses
ncAA Source
Endogenous - Biosynthesized
ncAA Availability
Can be synthesized in E coli expressing biosynthetic enzymes (see 2019 foundational paper) or purchased from commercial sources, such as Sigma-Aldrich catalog #81838
RS/tRNA Pair Usage Information
PraRS allows Pra to be incorporated into all Met positions all proteins in E. coli.
ncAA Synonyms
L-C-Propargylglycine
L-Propargylglycine
propargylglycine
Pra
(S)-2-Aminopent-4-ynoic acid
(2s)-2-aminopent-4-ynoic acid
4-Pentynoic acid, 2-amino-, (2S)-
L-2-Propynylglycine
(S)-2-Propargylglycine
(S)-2-Amino-4-pentynoic acid
(S)-Propargylglycine
(2S)-2-amino-4-Pentynoic acid
(S)-alpha-propargylglycine
L-Propargylglycine
propargylglycine
Pra
(S)-2-Aminopent-4-ynoic acid
(2s)-2-aminopent-4-ynoic acid
4-Pentynoic acid, 2-amino-, (2S)-
L-2-Propynylglycine
(S)-2-Propargylglycine
(S)-2-Amino-4-pentynoic acid
(S)-Propargylglycine
(2S)-2-amino-4-Pentynoic acid
(S)-alpha-propargylglycine
ChEBI ID
43797
PubChem Link