RS/tRNA Foundational Publication Support
Beyer, Jenna N, Parisa Hosseinzadeh, Ilana Gottfried-Lee, Elise M Van Fossen, Phillip Zhu, Riley M Bednar, Andrew Karplus, Ryan A Mehl, and Richard B Cooley. (2020) 2020. “Overcoming Near-Cognate Suppression In A Release Factor 1-Deficient Host With An Improved Nitro-Tyrosine Trna Synthetase.”. Journal Of Molecular Biology 432 (16): 4690-4704. doi:10.1016/j.jmb.2020.06.014.
RS/tRNA Protocols and Structural Information
Beyer, Jenna N, Parisa Hosseinzadeh, Ilana Gottfried-Lee, Elise M Van Fossen, Phillip Zhu, Riley M Bednar, Andrew Karplus, Ryan A Mehl, and Richard B Cooley. (2020) 2020. “Overcoming Near-Cognate Suppression In A Release Factor 1-Deficient Host With An Improved Nitro-Tyrosine Trna Synthetase.”. Journal Of Molecular Biology 432 (16): 4690-4704. doi:10.1016/j.jmb.2020.06.014.
RS/tRNA Usage Publications
Van Fossen, Elise M, Sonia Grutzius, Carl E Ruby, Dan Mourich V, Chris Cebra, Shay Bracha, Andrew Karplus, Richard B Cooley, and Ryan A Mehl. (2022) 2022. “Creating A Selective Nanobody Against 3-Nitrotyrosine Containing Proteins.”. Frontiers In Chemistry 10: 835229. doi:10.3389/fchem.2022.835229.
Zhu, Phillip, Kyle T Nguyen, Aidan B Estelle, Nikolai N Sluchanko, Ryan A Mehl, and Richard B Cooley. (2023) 2023. “Genetic Encoding Of 3-Nitro-Tyrosine Reveals The Impacts Of 14-3-3 Nitration On Client Binding And Dephosphorylation.”. Protein Science : A Publication Of The Protein Society 32 (3): e4574. doi:10.1002/pro.4574.
RS/tRNA Pair Development Year
2020
ncAA(s) Incorporated
3-nitro-L-tyrosine
ncAA Structure (png, jpg, jpeg)
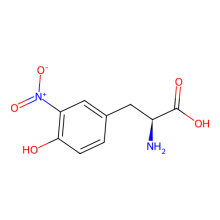
ncAA Utility
3-nitroTyr is a natural post-translational modification of proteins typically resulting from protein oxidation by peroxynitrite. It increases in many diseases and with age. Still unknown is if it can be reversed (by a denitrase) and if it has has a purposeful signaling role, or if it is only a modification associated with pathologies. It is known in some cases to contribute to disease pathology, but in most cases it is unknown how much it causes pathology vs. being a consequence of pathology. The ability to site specifically and quantitatively place 3-nitroTyr in positions in proteins that have been seen to occur in association with disease is a key tool to answer questions about its possible roles in normal physiology and disease.
RS Organism of Origin
Parent RS
RS Mutations
Y32H
H70T
D158H
I159A
L162R
H70T
D158H
I159A
L162R
tRNA Organism of Origin
Parent tRNA
tRNA Anticodon
CUA
Other tRNA Mutations
C17A
U17aG
U20C
G37A
U47G
U17aG
U20C
G37A
U47G
RS/tRNA Availability
on PDule plasmid as Addgene #174078
on pDule2 plasmid as Addgene #174079
on pDule2 plasmid as Addgene #174079
Used in what cell line?
RS/tRNA Additional Notes
This 3rd generation Mj RS/tRNA pair for 3-nitroTyr (3NY) outperforms earlier ones. Useful for full-fidelity incorporation of 3NY in truncation free RF1- strain. UP50 is ~25 microM and using 1 mM 3NY yields ~160 mg/L sfGFP(150) and about half as much dual-ncaa sfGFP(134/150). Also generated homogeneous 14-3-3(130) and calmodulin(99) at ~75 mg/L and 50 mg/L, respectively. A crystal structure with 3NY bound was also reported.