RS/tRNA Foundational Publication Support
Chin, Jason W, Ashton Cropp, Christopher Anderson, Mridul Mukherji, Zhiwen Zhang, and Peter G Schultz. (2003) 2003. “An Expanded Eukaryotic Genetic Code.”. Science (New York, N.y.) 301 (5635): 964-7.
Deiters, Alexander, Ashton Cropp, Mridul Mukherji, Jason W Chin, Christopher Anderson, and Peter G Schultz. (2003) 2003. “Adding Amino Acids With Novel Reactivity To The Genetic Code Of Saccharomyces Cerevisiae.”. Journal Of The American Chemical Society 125 (39): 11782-3.
Takimoto, Jeffrey K, Katrina Adams, Zheng Xiang, and Lei Wang. (2009) 2009. “Improving Orthogonal Trna-Synthetase Recognition For Efficient Unnatural Amino Acid Incorporation And Application In Mammalian Cells.”. Molecular Biosystems 5 (9): 931-4. doi:10.1039/b904228h.
Seidel, Lisa, Barbara Zarzycka, Saheem Zaidi, Vsevolod Katritch, and Irene Coin. (2017) 2017. “Structural Insight Into The Activation Of A Class B G-Protein-Coupled Receptor By Peptide Hormones In Live Human Cells.”. Elife 6. doi:10.7554/eLife.27711.
RS/tRNA Protocols and Structural Information
Serfling, R, and I Coin. (2016) 2016. “Incorporation Of Unnatural Amino Acids Into Proteins Expressed In Mammalian Cells.”. Methods In Enzymology 580: 89-107. doi:10.1016/bs.mie.2016.05.003.
RS/tRNA Usage Publications
Coin, Irene, Vsevolod Katritch, Tingting Sun, Zheng Xiang, Fai Siu, Michael Beyermann, Raymond C Stevens, and Lei Wang. (2013) 2013. “Genetically Encoded Chemical Probes In Cells Reveal The Binding Path Of Urocortin-I To Crf Class B Gpcr.”. Cell 155 (6): 1258-69. doi:10.1016/j.cell.2013.11.008.
Li, Pan, Jingxiang Li, and Haiyan Ren. (2022) 2022. “Capture And Mass Spectrometry Analysis Of Effector-Substrate Complexes Using Genetically Incorporated Photo-Crosslinkers In Host Cells.”. Star Protocols 3 (4): 101882. doi:10.1016/j.xpro.2022.101882.
Marino, Valerio, Wanchana Phromkrasae, Michele Bertacchi, Paul Cassini, Krittalak Chakrabandhu, Daniele Dell'Orco, and Michèle Studer. (2024) 2024. “Disrupted Protein Interaction Dynamics In A Genetic Neurodevelopmental Disorder Revealed By Structural Bioinformatics And Genetic Code Expansion.”. Protein Science : A Publication Of The Protein Society 33 (4): e4953. doi:10.1002/pro.4953.
Banerjee, Chiranjib, Brandon Wey-Hung Liauw, and Reza Vafabakhsh. (2025) 2025. “Direct Effect Of Membrane Environment On The Activation Of Mglur2 Revealed By Single-Molecule Fret.”. Structure (London, England : 1993) 33 (4): 718-727.e4. doi:10.1016/j.str.2025.01.011.
Sadler, Fredrik, Ning Ma, Michael Ritt, Yatharth Sharma, Nagarajan Vaidehi, and Sivaraj Sivaramakrishnan. (2023) 2023. “Autoregulation Of Gpcr Signalling Through The Third Intracellular Loop.”. Nature 615 (7953): 734-741. doi:10.1038/s41586-023-05789-z.
Liauw, Brandon Wey-Hung, Arash Foroutan, Michael R Schamber, Weifeng Lu, Hamid Samareh Afsari, and Reza Vafabakhsh. (2022) 2022. “Conformational Fingerprinting Of Allosteric Modulators In Metabotropic Glutamate Receptor 2.”. Elife 11. doi:10.7554/eLife.78982.
Schamber, Michael R, and Reza Vafabakhsh. (2022) 2022. “Mechanism Of Sensitivity Modulation In The Calcium-Sensing Receptor Via Electrostatic Tuning.”. Nature Communications 13 (1): 2194. doi:10.1038/s41467-022-29897-y.
Liauw, Brandon Wey-Hung, Hamid Samareh Afsari, and Reza Vafabakhsh. (2021) 2021. “Conformational Rearrangement During Activation Of A Metabotropic Glutamate Receptor.”. Nature Chemical Biology 17 (3): 291-297. doi:10.1038/s41589-020-00702-5.
Ye, Shixin, Morgane Riou, Stéphanie Carvalho, and Pierre Paoletti. (2013) 2013. “Expanding The Genetic Code In Xenopus Laevis Oocytes.”. Chembiochem : A European Journal Of Chemical Biology 14 (2): 230-5. doi:10.1002/cbic.201200515.
RS/tRNA Pair Development Year
2009
ncAA(s) Incorporated
p-azido-L-phenylalanine (pAzF)
ncAA Structure (png, jpg, jpeg)
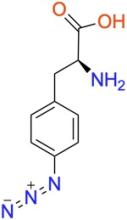
ncAA Utility
Used as a photocrosslinker, allowing for crosslinking and bioorthogonal click-chemistry ligation of proteins via strain-promoted cycloadditions (SPAAC) with suitably functionalized molecules.
RS Organism of Origin
Parent RS
RS Mutations
Y37L
D182S
F183M
L186A
D265R
D182S
F183M
L186A
D265R
tRNA Organism of Origin
Parent tRNA
tRNA Anticodon
CUA
Multiple tRNAs?
Shown in a 2002 paper (Sakamoto et al; PMID 12409460) the E. coli tRNA did not express well in mammalian cells and could be paired with the B. stearothermophilis suppressor tRNA-Tyr which expressed well. This is the tRNA used in plasmid available from Addgene.
RS/tRNA Availability
AddGene Plasmid #105829 encodes EAziRS and the B. stearothermophylus tRNA.
Used in what cell line?
RS/tRNA Additional Notes
2009 foundational paper added the D265R mutation to the RS reported as p-azidoPheRS-1 in 2003 foundational paper by Chin et al (2003) and as pAZ-EcRS1 by 2003 foundational paper by Dieters et al.. The D265R mutation in E coli Tyr RS improves its recognition of the CUA anticodon. The "E" in the RS name refers to that "enhanced" recognition. In HeLa cells it nearly doubled the fluorescence of produced GFP(39) with pAzPhe and the mutation also improved yields when put in RSs for three other ncAAs. In HEK293T cells. 1 mM pAzPhe incorporated into sites 87, 92 and 125 of GST showed increased yield and photocrosslinking of 30-100% of the protein.
The Addgene plasmid with this RS was developed in the 2017 foundational paper and seems to be the current go to system for mammalian cell studies with pAzPhe. It seems this same RS has also been called E2AziRS.
One oddity is that the 2009 paper reports the RS they studied as having an F183A mutation rather than the F183M mutation noted in the Addgene deposition. The F183A is present in p-azidoPheRS-3 from the 2003 papers.
The Addgene plasmid with this RS was developed in the 2017 foundational paper and seems to be the current go to system for mammalian cell studies with pAzPhe. It seems this same RS has also been called E2AziRS.
One oddity is that the 2009 paper reports the RS they studied as having an F183A mutation rather than the F183M mutation noted in the Addgene deposition. The F183A is present in p-azidoPheRS-3 from the 2003 papers.