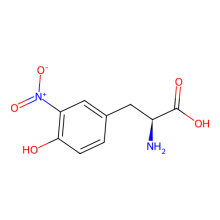
ncAA Structure (png, jpg, jpeg)
ncAA Foundational Publication Support
Neumann, Heinz, Jennifer L Hazen, John Weinstein, Ryan A Mehl, and Jason W Chin. (2008) 2008. “Genetically Encoding Protein Oxidative Damage.”. Journal Of The American Chemical Society 130 (12): 4028-33. doi:10.1021/ja710100d.
Cooley, Richard B, Jessica L Feldman, Camden M Driggers, Taylor A Bundy, Audrey L Stokes, Andrew Karplus, and Ryan A Mehl. (2014) 2014. “Structural Basis Of Improved Second-Generation 3-Nitro-Tyrosine Trna Synthetases.”. Biochemistry 53 (12): 1916-24. doi:10.1021/bi5001239.
Porter, Joseph J, Hyo Sang Jang, Elise M Van Fossen, Duy P Nguyen, Taylor S Willi, Richard B Cooley, and Ryan A Mehl. (2019) 2019. “Genetically Encoded Protein Tyrosine Nitration In Mammalian Cells.”. Acs Chemical Biology 14 (6): 1328-1336. doi:10.1021/acschembio.9b00371.
Gerding, Hanne, Christiaan Karreman, Andreas Daiber, Johannes Delp, Daniel Hammler, Martin Mex, Stefan Schildknecht, and Marcel Leist. (2019) 2019. “Reductive Modification Of Genetically Encoded 3-Nitrotyrosine Sites In Alpha Synuclein Expressed In E.coli.”. Redox Biology 26: 101251. doi:10.1016/j.redox.2019.101251.
Beyer, Jenna N, Parisa Hosseinzadeh, Ilana Gottfried-Lee, Elise M Van Fossen, Phillip Zhu, Riley M Bednar, Andrew Karplus, Ryan A Mehl, and Richard B Cooley. (2020) 2020. “Overcoming Near-Cognate Suppression In A Release Factor 1-Deficient Host With An Improved Nitro-Tyrosine Trna Synthetase.”. Journal Of Molecular Biology 432 (16): 4690-4704. doi:10.1016/j.jmb.2020.06.014.
Avila-Crump, Savanna, Marcus L Hemshorn, Chloe M Jones, Lea Mbengi, Kyle Meyer, Joshua A Griffis, Subhashis Jana, et al. (2022) 2022. “Generating Efficient Pyrrolysyl-Trna Synthetases For Structurally Diverse Non-Canonical Amino Acids.”. Acs Chemical Biology 17 (12): 3458-3469. doi:10.1021/acschembio.2c00639.
ncAA Usage Publications
Rauch, Benjamin J, Joseph J Porter, Ryan A Mehl, and John J Perona. (2016) 2016. “Improved Incorporation Of Noncanonical Amino Acids By An Engineered Trna(Tyr) Suppressor.”. Biochemistry 55 (3): 618-28. doi:10.1021/acs.biochem.5b01185.
Gerding, Hanne, Christiaan Karreman, Andreas Daiber, Johannes Delp, Daniel Hammler, Martin Mex, Stefan Schildknecht, and Marcel Leist. (2019) 2019. “Reductive Modification Of Genetically Encoded 3-Nitrotyrosine Sites In Alpha Synuclein Expressed In E.coli.”. Redox Biology 26: 101251. doi:10.1016/j.redox.2019.101251.
Zheng, Zhaopeng, Xuzhen Guo, Minling Yu, Xiaoyan Wang, Hongguang Lu, Fahui Li, and Jiangyun Wang. (2020) 2020. “Identification Of Human Ido1 Enzyme Activity By Using Genetically Encoded Nitrotyrosine.”. Chembiochem : A European Journal Of Chemical Biology 21 (11): 1593-1596. doi:10.1002/cbic.201900735.
ncAA Utility
3-nitroTyr is a natural post-translational modification of proteins typically resulting from protein oxidation by peroxynitrite. It increases in many diseases and with age. Still unknown is if it can be reversed (by a denitrase) and if it has has a purposeful signaling role, or if it is only a modification associated with pathologies. It is known in some cases to contribute to disease pathology, but in most cases it is unknown how much it causes pathology vs. being a consequence of pathology. The ability to site specifically and quantitatively place 3-nitroTyr in positions in proteins that have been seen to occur in association with disease is a key tool to answer questions about its possible roles in normal physiology and disease.
ncAA Source
Exogenous - Purchased
ncAA Availability
Can be purchased from https://www.sigmaaldrich.com/US/en/product/sigma/n7389
RS/tRNA Pair Usage Information
For work in E. coli, the best proven RS to use is the "3-nitroTyr(A7) RS/tRNA(CUA)" which gives high-fidelity incorporation even in truncation free RF- B95 cells. It is a 3rd generation RS, superceeding the "RS8" and "5B" RSs. For work in mammalian cells, the road-tested RS is "A7-nitroTyr-MbPylRS". The "F4" RS incorporated 3NY in E. coli, but not in mammalian cells. The D9 RS from M alvus (and some other RSs reported in that study) appears to work as well in E coli as the M barkeri A7 RS, but the selected M alvus RSs have not been tested in mammalian cells. Both the "3-nitroTyr(A7) RS/tRNA(CUA)" and the "A7-nitroTyr-MbPylRS" are available from Addgene.
RS/tRNA Pair(s)
ncAA Synonyms
3-Nitro-L-tyrosine
621-44-3
H-Tyr(3-NO2)-OH
3-Nitrotyrosine
L-3-Nitrotyrosine
nitrotyrosine
nitroTyr
3-NT
3-NY
3NY
621-44-3
H-Tyr(3-NO2)-OH
3-Nitrotyrosine
L-3-Nitrotyrosine
nitrotyrosine
nitroTyr
3-NT
3-NY
3NY
ChEBI ID
44454
PubChem Link
PDB Code
NIY
ncAA Additional Notes
2019 foundational paper noted that within E. coli cells 3NY on proteins can be reduced to 3-amino-Tyr, so this is something to watch out for.